Tau Uptake and Spread via LRP1
How does pathological tau move from cell to cell?
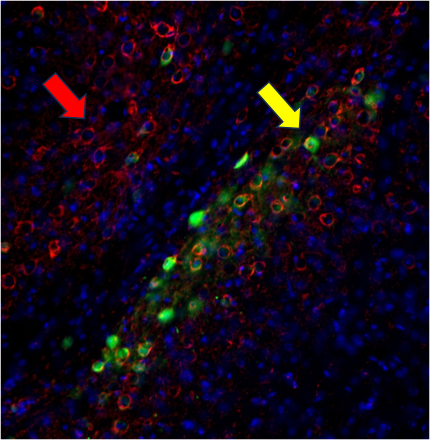
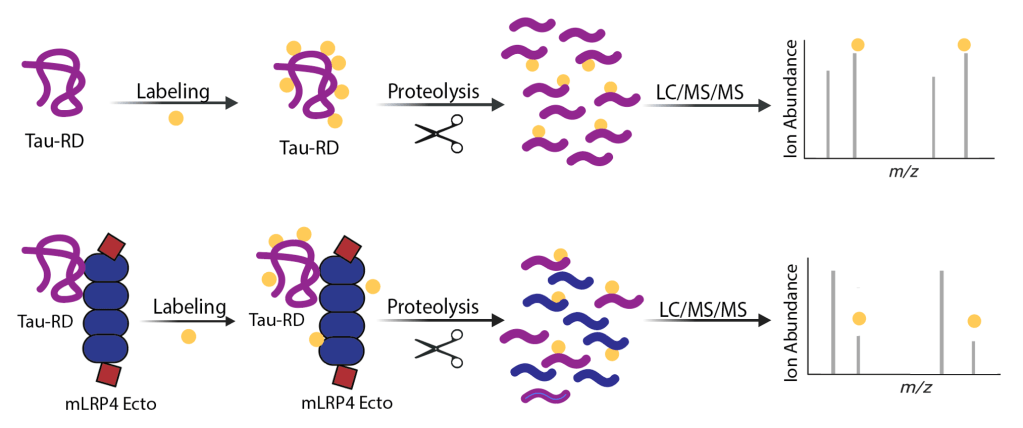
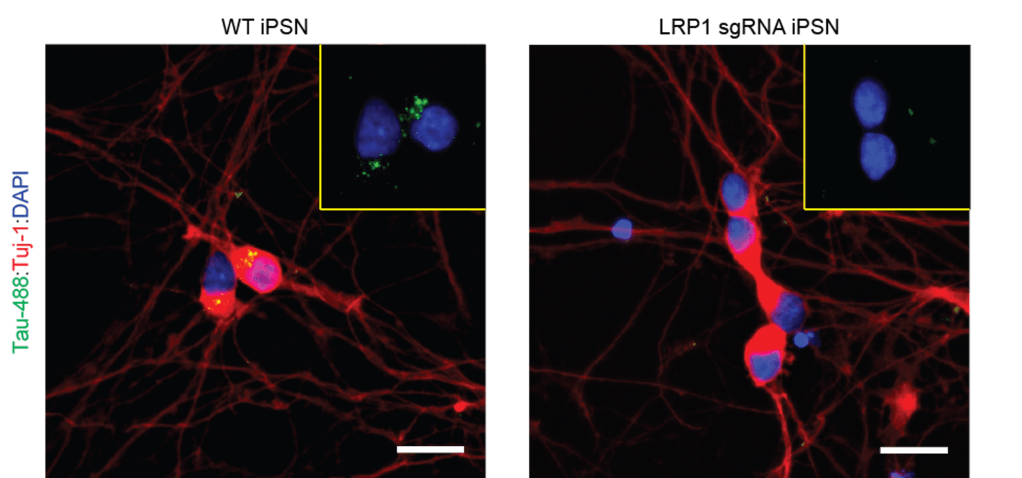
Tau aggregates propagate through the brain in a hierarchical pattern that tracks with cognitive decline, yet the molecular features that govern this spread are only beginning to be defined. Building on our discovery that LRP1 is a master regulator of tau uptake, we are dissecting the tau-LRP1 interaction at multiple scales. We use biochemical and biophysical approaches to map the tau-LRP1 interface and probe how post-translational modifications tune interaction. We examine how LRP1 functions in diverse cellular environments and are mapping the downstream consequences of tau-LRP1 binding. In parallel, we are mining chemical space using high-throughput and DNA-encoded library screening approaches to find molecules that disrupt the complex and block tau spread.
Tau Aggregation and Degradation
What turns soluble tau into a pathogenic aggregate — and how do cells try to stop it?
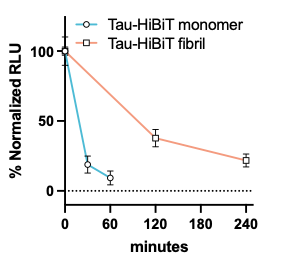
Tau is an abundant and highly soluble protein, yet in disease it transitions into stable aggregates. To understand the aggregation process better we’ve developed tau constructs and models to track aggregation kinetics in vitro, in cells, and in vivo. To understand how cells process tau, we’ve developed quantitative tau degradation platforms to map tau turnover. Together, these tools illuminate the pathways cells use to manage tau, and the points at which they fail.
Cell-Type-Specific Contributions to Tauopathy
Which cells in the brain drive tau spread, and which protect against it?
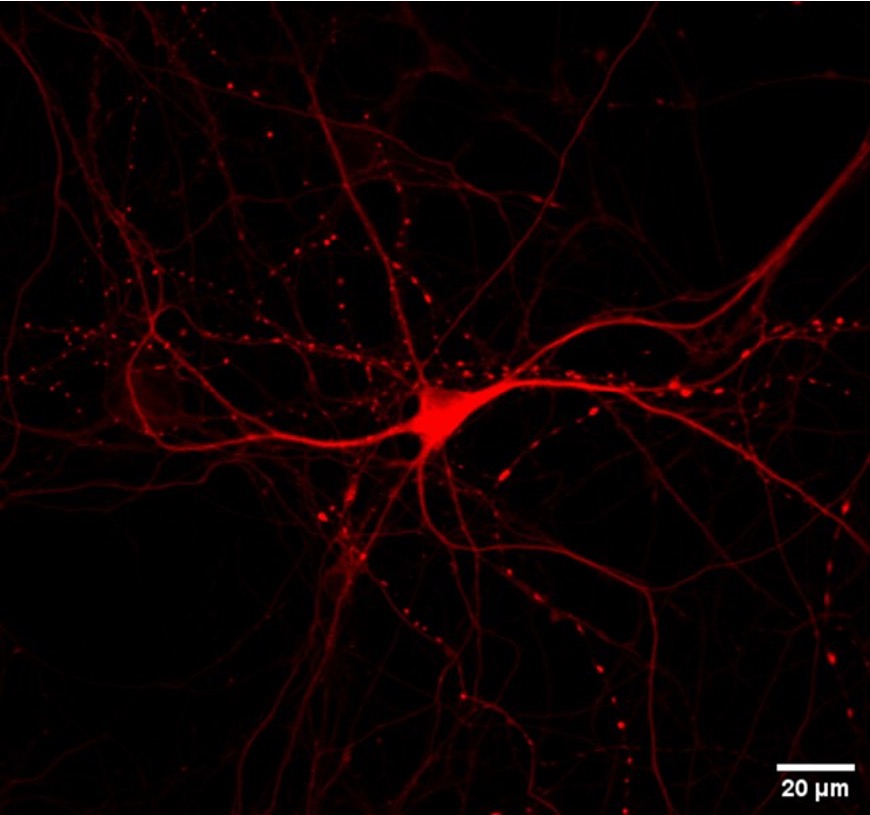
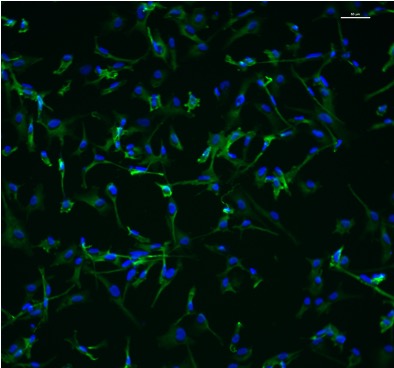
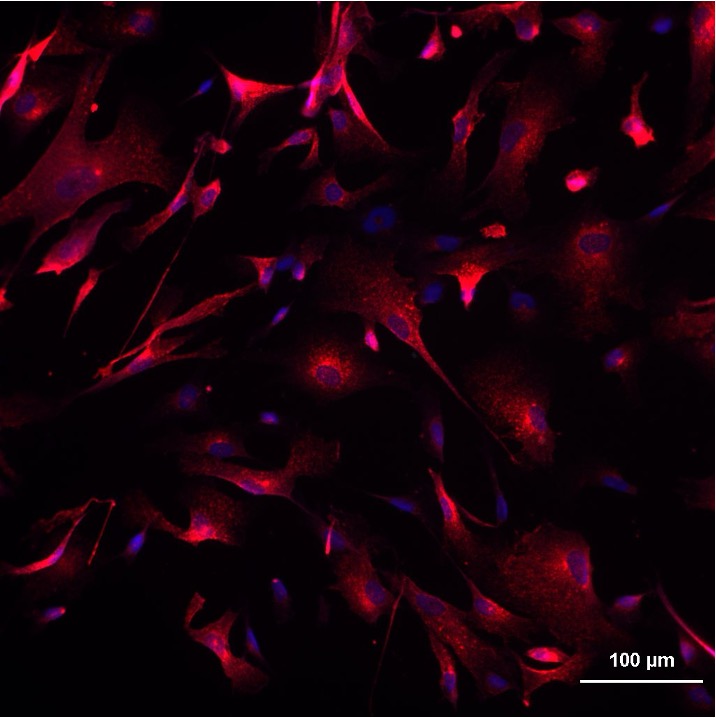
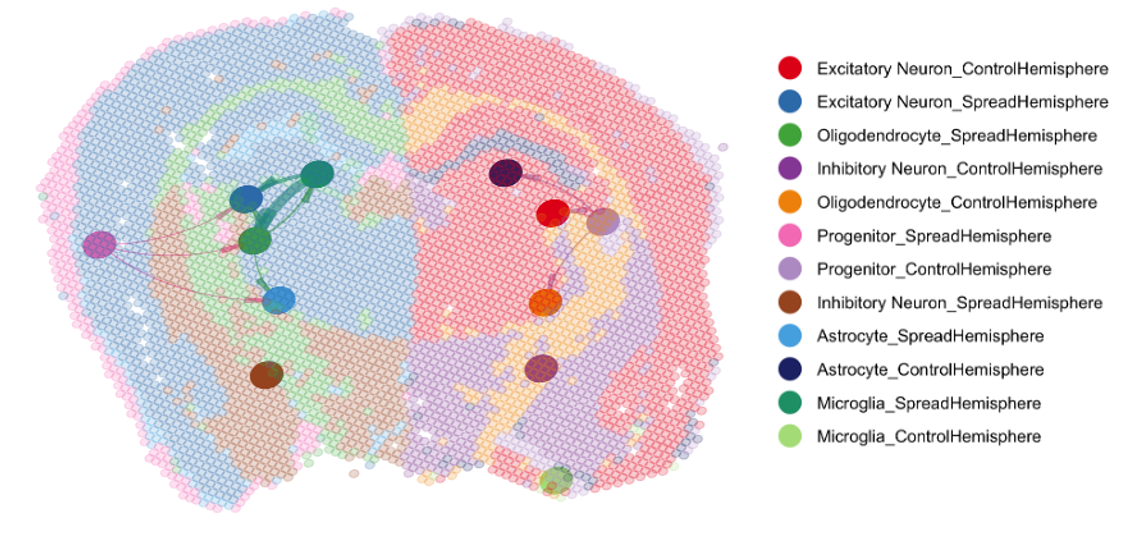
Most of our mechanistic understanding of tau aggregation comes from model cell lines (HEK293T) or neuron-centric systems. However, the brain is a complex tissue and different cell types may experience and propagate tau pathology in fundamentally different ways. We are building tools to study tau in this fuller context. Utilizing adeno-associated virus (AAV) we have developed sensors to monitor aggregation in defined neuronal and glial populations, while single-cell RNA sequencing, proteomics, and spatial transcriptomics we aim to map how cell identity shapes tau quality control.